We recently added PDF processing and de-identification capabilities to CLEHR. This was driven directly by what our customers needed: sponsors told us that without access to PDF attachments in the EHR, the promise of automated eSourcing was incomplete. CRCs could see information that the data pipeline could not. And in the process of building this capability, we learned things about what is actually in the clinical record—the format landscape, the hidden metadata, the redundancy between document types—that we think are worth sharing, because they affect anyone working with EHR data for research or clinical trials.
This post covers what we built, the practical and technical surprises along the way, and the use cases we see as most significant.
Background: From Structured Data to Documents
CLEHR started as a structured data problem. The core use case was eliminating the manual transcription burden of EHR-to-EDC workflows—getting structured patient data from Epic and other EHR systems into sponsor EDCs without a CRC manually entering values field by field. Getting that right required building de-identification infrastructure first: a compliant, auditable pipeline for handling PHI before any data leaves the health system. Once that foundation was in place, it became clear that de-identification capability opened a broader set of use cases. Free-text clinical notes were the natural next step—applying the same pipeline to the unstructured data that structured queries cannot reach.
PDFs are the next layer in that progression. They are not part of the EHR’s native data model—they are attachments, documents that arrive from outside the system or are generated by processes that do not write back to structured fields. In many cases they carry some of the most precise, most specialized clinical data in the entire record. And like free-text notes before them, they are completely opaque to any tool that does not read the document itself.
Nobody Really Knows What’s in the Record
Here is the structural reality that underlies everything in this post: nobody actually knows what’s in the FHIR record. Not the clinicians. Not the research departments. Not the IT teams managing the infrastructure. The data is there—someone put it in, and someone can open it and read it—but no one has systematically analyzed what’s actually present across the full breadth of the record, including its attachments.
This is not a FHIR-specific problem. It is an enterprise data management problem in the context of large hospitals running Epic or other major EHR systems. Data arrives through structured lab interfaces, through free-text notes, through HTML documents with inline images, and through PDF attachments. It gets stored. But the relationship between what was documented and where it ended up is not something anyone can easily map.
When a clinician orders a test through the EHR—a standard CBC, a metabolic panel—the result returns through the lab interface and populates a structured field. It is queryable. You can find every patient with a hemoglobin below 8, or every patient whose creatinine crossed a threshold during a study period.
But a large category of specialized testing does not work this way. Genetic panels ordered through external laboratories, detailed metabolic studies from reference labs, results from testing platforms that have their own patient-facing portals—these come back as PDFs. The clinician downloads the report, uploads it into Epic as an attachment, and it enters the record as a document. A physician can open it and read it. A CRC doing manual chart review can find it. But no FHIR query, no cohort builder, no structured analytics platform can tell you what is in it.
I had a conversation at a recent conference with an endocrinologist who specializes in pediatric obesity. She orders genetic panels regularly—testing for pathogenic variants in genes like MC4R, PCSK1, and LEPR that define specific monogenic obesity subtypes. The labs return those results through an external portal. She downloads them and uploads them into Epic. The data is in the record. But she cannot identify her patients by gene variant. She cannot filter her panel by mutation type. She cannot ask whether her patients with MC4R variants respond differently to GLP-1 agonists than those with polygenic obesity. That analysis—which would be straightforward if the genetic data were structured—is effectively impossible because the results live in PDFs.
This pattern repeats across specialties: the more specialized the test, the more likely the result comes back as a document rather than populating a structured field. Outside institution reports, reference laboratory panels, subspecialty consultation summaries from non-affiliated practices—all of these commonly arrive as attachments. The information is present in the record but opaque to any tool that does not read the document.
What’s Actually in There: PDFs, HTML, and the Formats You Didn’t Expect
When we started working with the document layer of the FHIR record, one of the first things we discovered is that the format landscape is more varied than most people assume. In the FHIR data model, clinical documents can live in two places—DocumentReference resources and DiagnosticReport narratives—and the content can arrive as plain text, rich text, HTML, or PDF. What you get depends on the source system, the site configuration, and sometimes the specific type of result.
The HTML finding surprised us, and it surprises most people we mention it to. Clinical documents stored as HTML are not just formatted text. They are often self-contained HTML documents with inline embedded images—ECG screenshots, facility logos, digital signatures. The images are base64-encoded directly in the HTML, not referenced from remote URLs. Everything is in the document. This means that processing these documents is not simply a text extraction problem—you have to handle the embedded media as well, both for de-identification and for any downstream use of the content.
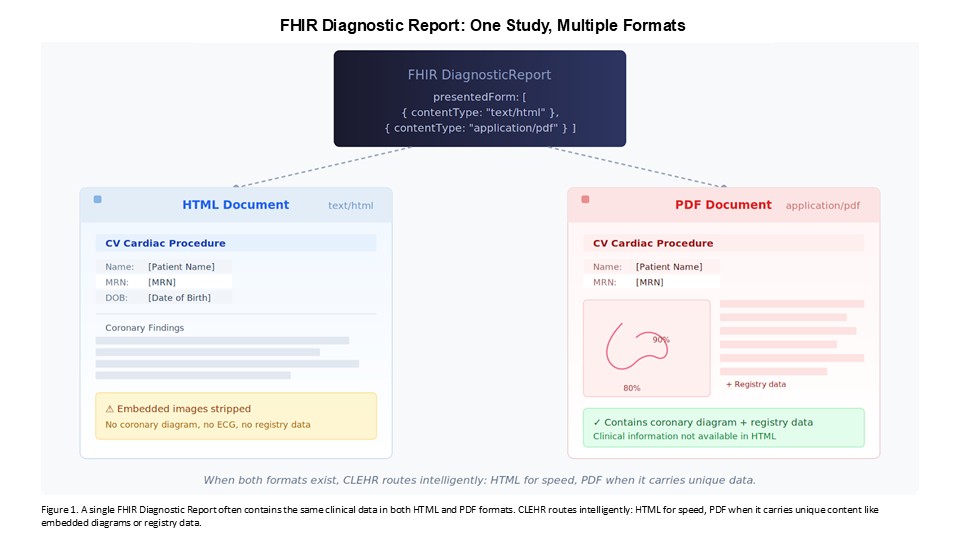
In many cases, the same clinical report exists in multiple formats within the record—an HTML version and a PDF version of the same study, linked to the same diagnostic order. This redundancy creates an important routing decision. HTML is substantially easier and faster to process: the text is directly parseable, and our pipeline handles it the same way it handles other unstructured text in the system. PDF requires OCR, layout analysis, and additional compute. When both formats are available, we take the HTML. We process the PDF only when it is the sole available format or when the PDF contains information—graphical content, embedded tables, registry data—that did not make it into the HTML representation.
That routing logic matters operationally. It reduces processing overhead for the sites and for sponsors, and it means the decision about when to incur the cost of PDF processing can be made intelligently—per study, per document type, per clinical context.gr

What We Built
The CLEHR PDF pipeline has three components: extraction, de-identification, and context preservation. Each required design decisions specific to clinical documents, not generic document processing.
Extraction
We use a combination of Tesseract OCR and AWS Textract for text extraction, selected based on document characteristics. The choice of extraction approach matters more for clinical documents than it might for general text because the layout carries meaning. A genetic report organizes findings in a structured table—gene name, variant identified, pathogenicity classification, clinical interpretation. A cardiology procedure note might have measurements embedded in a narrative with specific anatomical references. Extracting these as undifferentiated character strings loses the relationships between fields that give the data its clinical meaning.
Our extraction layer preserves document structure: tables are extracted with their spatial relationships intact, section headers are recognized and preserved, and the associations between labels and values are maintained in the output. The goal is not to produce a flat text dump but to produce something that retains enough of the original document’s organization to support downstream analysis and de-identification.
De-identification: Hidden in Plain Sight
Once extracted, the text goes through CLEHR’s de-identification pipeline using Tonic.ai’s redaction tooling, consistent with the approach we use across the platform for unstructured EHR data. This handles the standard PHI categories—names, dates, geographic identifiers, contact information, identifiers—but clinical documents from outside institutions often carry additional identifying material that generic de-identification tools do not handle well: provider names, facility names, and order numbers from external laboratory systems. Our pipeline has been extended to address these document-specific categories.
For PDFs specifically, we use a synthesis-based approach rather than simple blackout redaction. Instead of replacing a patient name with a black bar—which signals to any reader exactly where PHI was present—our pipeline replaces identifying text with synthetic but realistic substitutions. A real patient name is replaced with a synthetic name. The replacement is rendered in the matching font, at the correct size, in the correct position within the document. The result is a PDF that reads naturally, where the de-identified elements are indistinguishable from the original text.
This approach, sometimes called “hidden in plain sight,” provides an important additional layer of protection. With traditional blackout redaction, if a name is redacted you know it was real PHI—any leak is immediately identifiable as a breach. With synthesis, there is plausible deniability: every name in the document could be synthetic, and there is no visual marker distinguishing original text from replacements. This is meaningfully harder to implement for PDFs than for plain text, because the font matching, character spacing, and layout fidelity have to be precise—otherwise the synthetic text stands out and the protection breaks down.

The de-identification is also configurable per study. Date handling, for example, can follow different policies: consistent date shifting across a patient’s record, age-aware shifting that avoids creating impossible timelines for pediatric populations, or preservation of age at the expense of exact dates. Gender-consistent name replacement, category-specific redaction policies—these are parameters that can be tuned to the regulatory and analytic requirements of each engagement.
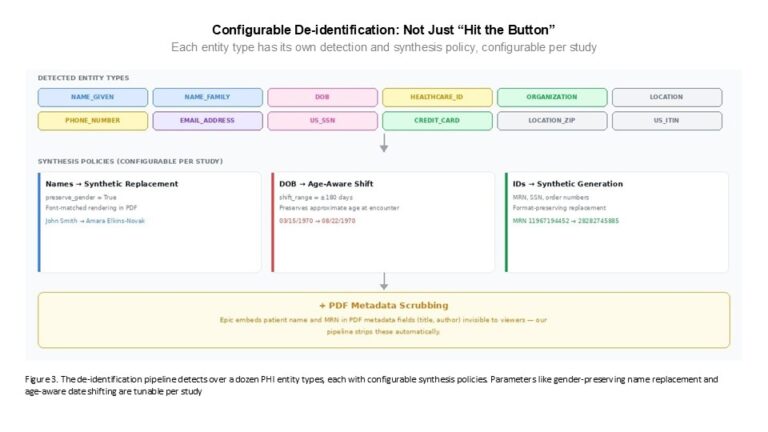
Metadata: The PHI You Don’t See
One practical finding that has significant compliance implications: Epic embeds patient identifying information—including the patient name and medical record number—in the metadata of PDFs it generates. This metadata is not visible when you open the document and read it. It is not visible in a standard viewer. But anyone with a PDF metadata reader can extract it, and if a de-identification pipeline processes only the visible text content of the document, that PHI passes through undetected.
Our pipeline scrubs PDF metadata as part of the standard processing workflow. This is the kind of detail that is easy to overlook and consequential to miss.
The FHIR Pipeline Problem
One practical implication of PDF processing that is easy to overlook: PDFs transit through FHIR pipelines even for organizations that are not explicitly trying to process them for analytics. The FHIR DocumentReference resource is a standard mechanism for transmitting clinical documents, and when sites use FHIR for EHR-to-EDC workflows, PDFs arrive in the data stream regardless of whether anyone asked for them. That means even if a sponsor has no interest in analyzing PDF content, they need a decision about what to do with those documents from a compliance and de-identification standpoint. Our pipeline addresses that as well.
Why Sponsors Need This: The CRC Parity Problem
The driver for PDF processing in clinical trials is straightforward, even if the implications are broad: sponsors need to know that the data they receive through eSourcing workflows gives them access to everything a research coordinator behind the firewall has access to. If it does not, the entire value proposition of direct EHR integration is compromised.

Consider a simple example. A patient undergoes an interventional cardiology procedure. The procedure report, stored as a PDF attachment, documents that the patient experienced a hematoma at the access site. The EDC for the study has a field asking whether the patient experienced a hematoma during the procedure. A CRC reading the chart checks yes. But if the automated data capture workflow only processes structured data and free-text notes, and the hematoma was documented in the PDF procedure report, the eSourcing system misses it. The data is incomplete—not because it was absent from the record, but because it was in a format the system did not process.
This is the core problem. As long as there is a category of clinical documentation that automated systems cannot reach, there is uncertainty about completeness. Even if 90% of relevant information is in structured fields and notes, the remaining 10% in PDFs creates a coverage gap that the sponsor cannot quantify—because you don’t know what you don’t know.
There is also a practical cost dimension. Even when CRCs are handling data entry, sponsors currently ask them to download PDFs from Epic, upload them to third-party source document tools, and transmit them through separate channels for review. The third-party tools attempt redaction with variable quality. The workflow is manual, labor-intensive, and detached from the rest of the clinical data capture process. CRCs are being asked to manage a parallel document pipeline on top of their existing workload—and compliance is inconsistent, because the process depends on each coordinator remembering to upload every relevant PDF for every patient.
Sponsors know this is painful for sites. Things that are painful for sites create reluctance to participate in studies, drive up costs, and degrade data quality. When a site has to choose between a study that requires manual PDF upload workflows and one that does not, the decision is easy—and it is not in the sponsor’s favor.
PDF processing within CLEHR eliminates this parallel workflow. Documents flow through the same pipeline as structured data and notes, de-identified and delivered under the same governance framework. The CRC does not need to manually handle source documents. The sponsor receives the complete documentary record—structured fields, notes, and PDFs—through a unified system. One pipeline to monitor, one pipeline to validate, one pipeline for the site to manage.
Applications for Real-World Evidence
The primary RWE value of PDF processing is cohort enrichment—the ability to characterize patients using data that structured queries cannot reach. The cases where this matters most tend to be the cases where specialized subpopulation identification is most important:
- Genetic driver identification. Finding patients with specific pathogenic variants from outside laboratory reports embedded in the EHR. The endocrinologist’s use case described above—identifying MC4R or PCSK1 patients for comparative effectiveness analysis—is a representative example. The same pattern applies across oncology, rare disease programs, and any indication where genetic stratification matters for study design or outcomes analysis.
- Outside institution results. Patients in specialty care frequently have relevant testing and consultations at institutions other than the one where a study is being conducted. Those results arrive as documents. Ignoring outside results means working with an incomplete clinical picture.
- Reference laboratory panels. Specialized testing platforms—comprehensive metabolic panels from reference labs, detailed endocrine assays, allergy and immunology panels—frequently return results as formatted reports rather than discrete structured values. This is particularly common in pediatric subspecialties and rare disease care.
- Procedure and imaging reads from outside facilities. When patients receive imaging or procedures at affiliated or outside institutions that transmit reports rather than structured data, the clinical findings live in the PDF. For cardiovascular, oncology, and musculoskeletal indications, this can include quantitative measurements and severity assessments central to study endpoints.
- Embedded registry data. In some clinical contexts—interventional cardiology in particular—PDFs contain structured data that was originally assembled for clinical registries. These registry submissions represent carefully curated, standards-formatted data about procedures, outcomes, and device usage. When this data is embedded in the PDF, it is a high-value extraction target: essentially pre-structured information that would otherwise require significant manual effort to reconstruct.
Across our European research network—including our partnerships through HUGO, APHP, and our Italian hospital partners via Biomeris—PDF processing also addresses a practical interoperability reality. Many institutions in federated network environments transmit clinical documents as PDF attachments rather than structured FHIR resources, particularly for historical records and outside results. Structured queries against those networks return an incomplete picture without PDF processing.
Applications for Clinical Trials
In clinical trial settings, PDFs create a specific challenge for source data workflows. The issue is not just that data is inaccessible for analytics—it is that the most careful and complete source data capture processes can still miss information that is present in the record but embedded in an attachment.
As sponsors move toward automated EHR-to-EDC workflows, the assumption is often that what comes through FHIR is what there is. In practice, a clinically meaningful result—a genetic test that determines eligibility, a specialty evaluation documenting a key baseline characteristic, a reference lab panel capturing a study endpoint—may be present in the chart as a PDF attachment and absent from the structured data stream entirely. The CRC knows it is there. The monitor can see it during a site visit. But the automated data capture missed it.
This creates the source data integrity concern that EHR-to-EDC is supposed to eliminate. The point of direct EHR integration is to get data that is accurate from the source, with no transcription error and no gaps from manual abstraction. When key values live in PDFs that the integration cannot process, that promise is only partially delivered.
PDF processing within CLEHR addresses this by making document-embedded values available for EDC population alongside the structured data, while preserving the original PDF as the source documentation. This is not a workaround—it reflects the actual information architecture of how clinical data flows at sites. The record includes structured fields and it includes documents, and a complete eSourcing solution needs to handle both.
There is also a screening and eligibility dimension. Studies that require outside test results, specialty evaluations, or genetic data as part of eligibility determination currently rely on manual abstraction by study staff. Automated PDF extraction can support eligibility screening against criteria that reference values embedded in attachments, reducing manual burden and improving consistency across sites.
Looking ahead, we are building toward a system that can either automatically validate what the CRC entered—essentially automated source data verification—or perform the data extraction itself, reducing the need for SDV because the system processed the same source material the CRC would have reviewed. Either path requires the ability to process PDFs, because the source data lives there.
Non-Text Content: The Next Frontier
Not everything in a PDF is text. Clinical procedure reports—particularly in interventional cardiology—routinely embed diagrams, annotated images, and graphical representations that carry clinical information with no corresponding text description. In reviewing PCI procedure reports, we have observed diagrams indicating the location and severity of coronary occlusions, with spatial annotations documenting which vessels are affected and the degree of stenosis. These are essentially visual encodings of clinical data—like the numbered tooth charts dentists use, but for coronary anatomy—and they exist only in the image.
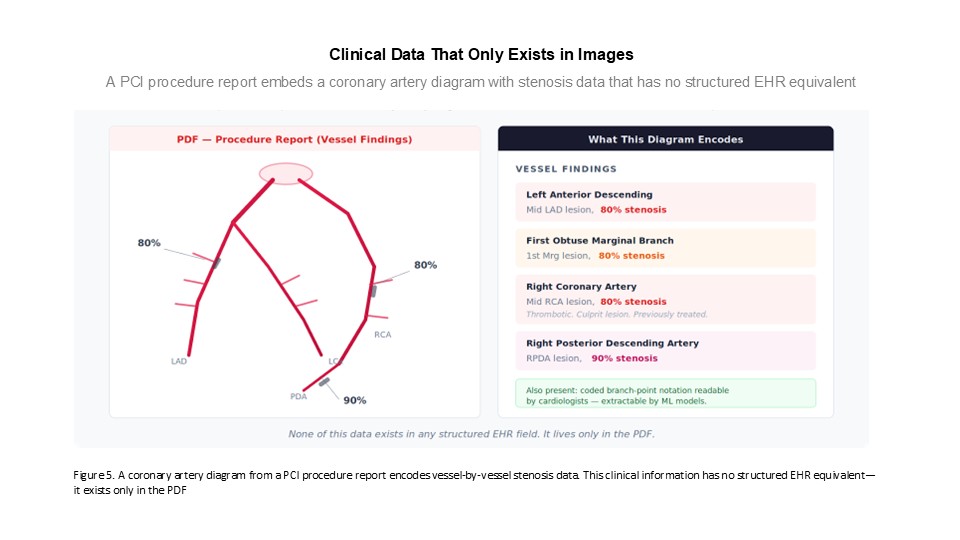
In some cases, there are also textual representations of this same anatomical data: coded branch-point notation systems that cardiologists can read directly. Both the graphical and textual forms live in the PDF, and both are invisible to any system that processes only structured EHR data.
Similarly, embedded HTML documents within the EHR can contain ECG screenshots, waveform images, and other graphical clinical data. These are currently outside the scope of our text extraction and de-identification pipeline, but they represent a real data access problem: there is no text-equivalent for an ECG tracing or a coronary artery diagram.
Our current pipeline handles text extraction and de-identification. Extending it to image extraction and redaction within PDF documents is the next phase of development—both to enable research use of embedded graphical data and to ensure compliant handling of images that may carry protected health information, such as photographs or annotated diagrams that include patient-identifiable elements.
What We Are Looking For
PDF processing within CLEHR is available now. If your organization has document-processing challenges in clinical research workflows, we can work with you today. The cases that are most interesting to us right now:
- Health systems that have significant clinical data locked in PDF attachments and want to make it accessible for research or quality programs, particularly in specialty areas like genetics, cardiology, and oncology where outside results are common.
- Pharma sponsors running RWE studies or hybrid trial designs where outside results and specialist documentation are part of the evidence base and structured data alone does not fully characterize the population.
- Clinical sites and CROs where PDF-based source data creates gaps in EHR-to-EDC workflows—either because key values are missing from structured FHIR output or because document-embedded data is not being captured in the EDC at all.
- Organizations with complex de-identification requirements for clinical documents, particularly where standard PHI categories do not cover all identifying elements present in outside institution reports.
We are also building the image extraction and redaction capability for the next phase. If your organization has specific use cases involving embedded images or graphical content in clinical PDFs—procedure reports, pathology slides, annotated imaging reads—we would be interested in understanding what that looks like in practice.
This expertise also extends beyond the CLEHR eSourcing context. If your organization is running real-world evidence studies and PDFs are a data access problem, we have the infrastructure and experience to work through your specific PDF challenge—whether that means integrating with our platform or applying these capabilities in a standalone engagement.
If any of this connects to challenges you are working through, reach out to the Graticule team at info@graticule.life.
CLEHR is Graticule’s NLP/AI platform for unstructured EHR data extraction and de-identification, supporting EHR-to-EDC workflows, real-world evidence studies, and federated research networks across the US and Europe.